Transcriptional recording with Record-seq
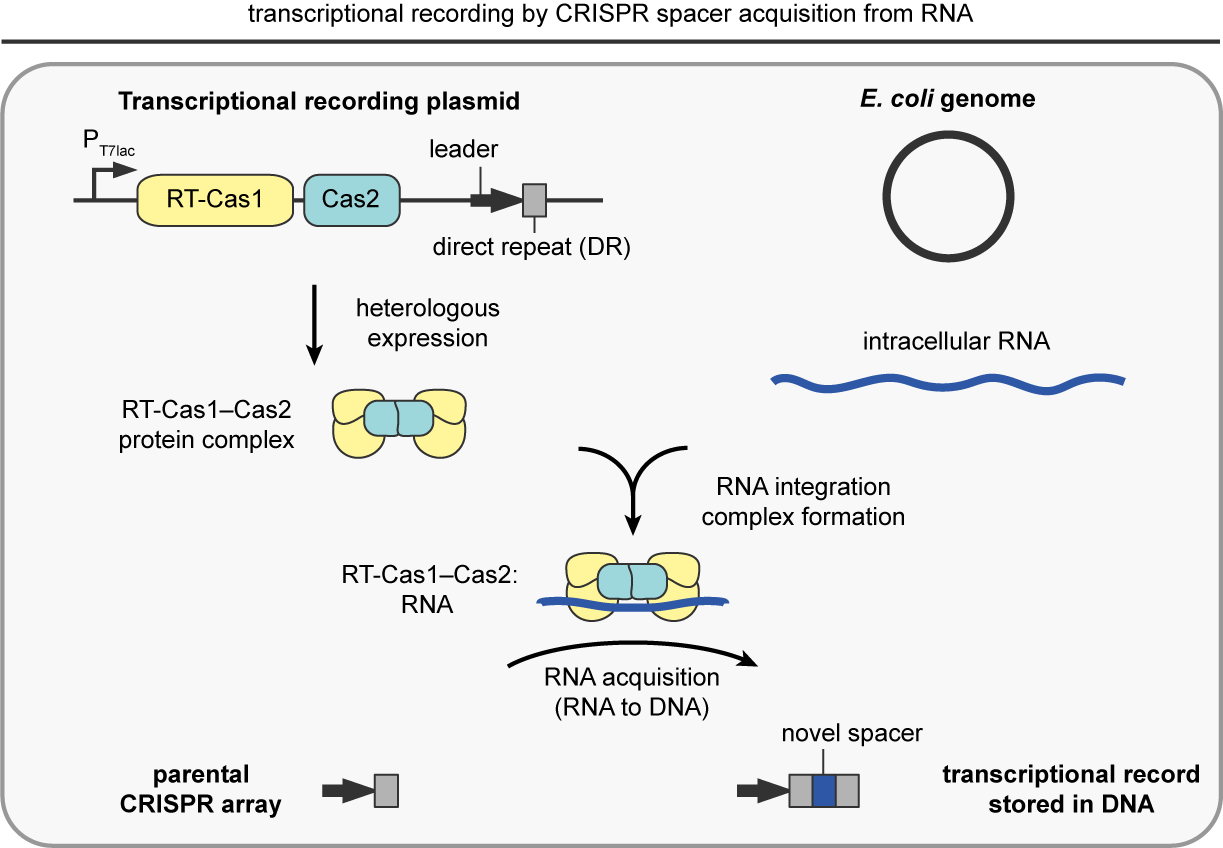
Transcriptional recording by CRISPR spacer acquisition from RNA
Microorganisms evolved to be experts in adapting and surviving in profoundly diverse environments – water, soil, air, and seemingly all surfaces of the human body – and their gene expression patterns reflect the environments in which they live. For example, in the inflamed human intestine, commensal bacteria adapt to a harsh, free radical-rich environment by coordinating the expression of specific scavengers and detoxifying agents. This presents a promising clinical opportunity where microbial gene expression patterns could be leveraged to assess the intestinal environment for non-invasive diagnostics and as a basis for devising individualized clinical interventions. Unfortunately, however, microbial gene expression information does not persist throughout time and transit of the gastrointestinal tract and is therefore inaccessible without invasive collection.
To overcome this challenge, my laboratory created “transcriptional recording”, a technology that enables engineered bacteria to sense, remember, and report on diverse intracellular and extracellular environments. We accomplished this by hijacking the natural process of CRISPR spacer acquisition from RNA, wherein short segments of intracellular RNAs, derived from endogenous transcripts or invading genetic elements, are reverse transcribed into DNA and integrated into a CRISPR array via the action of two CRISPR-associated proteins. Inside of living cells, the transcriptional recording machinery continuously captures RNAs proportional to their abundance and permanently stores them within DNA. This process is also iterative, whereby new RNAs are inserted in front of old RNAs within the CRISPR array – like a fossil record storing ordered, historical information. Upon sequencing the CRISPR arrays of a population of many cells, the RNA-derived segments can be aligned to the genome of the bacterial host and used to quantitatively reveal a transcriptional history over time (Schmidt, Nature, 2018; Tanna, Nature Protocols, 2020). Over the past several years we have demonstrated that transcriptional recording sentinel cells can traverse the gastrointestinal tract of animals and report on complex and multi-faceted environments critical for diagnostics, including pathogenic infections, alterations in diet, and pathological environments such as inflammation (Schmidt et al., Science, 2022). In sum, living microbial sentinel cells are capable of continuously recording the molecular and chemical properties of their environment – opening up radically new avenues for interrogating our world and improving human health.
CRISPR spacer aquisition from RNA
Downloads and links
Record-seq experimental protocol, including Selective amplification of ExpaNdEd Crispr Arrays (SENECA)
external page Tanna et al. Nature Protocols, 2020
Record-seq code
external page Platt Lab Transcriptional Recording GitHub
Deep sequencing data (Sequencing Read Archive)
external page Schmidt et al., Nature 2018
external page Tanna et al., Nature Protocols 2020
external page Schmidt et al., Science 2022
Plasmids (Addgene)
external page Schmidt et al., Nature 2018
external page Schmidt et al., Science 2022
Further reading
Schmidt et al., Nature 2018
- external page The Scientist
- external page Nature
- external page Nature Biotechnology
- external page Nature Methods
- external page Nature Chemical Biology
- ETH News
- external page The Science Breaker
Schmidt et al., Science 2022
- external page Science
- ETH News
- external page Insel News
- external page BRCCH News
- external page GEN News